The Kruskal algorithm is a greedy algorithm used to find a Minimum Spanning Tree (MST) in a graph with weights on the edges. A Minimum Spanning Tree (MST) of a graph is a subset of the edges that connects all the nodes of the graph such that the sum of the weights of the edges is minimal.
[wpda_org_chart tree_id=44 theme_id=50]
Kruskal’s algorithm
Kruskal’s algorithm was proposed by Joseph Kruskal in 1956. Joseph Kruskal is an American mathematician and computer scientist, known for his contributions to the fields of algorithms, graph theory, and computer networks. The algorithm was presented in a paper titled “On the Shortest Spanning Subtree of a Graph and the Traveling Salesman Problem” published in the journal Proceedings of the American Mathematical Society.
Kruskal’s algorithm is a greedy algorithm that is based on selecting the edges of the graph in ascending order with respect to the weights. The goal is to build a minimal spanning tree by edge selection such that no loops are formed. The algorithm uses the Union-Find (or Disjoint Set) data structure to manage the connected components of the graph and to avoid the formation of loops during the construction of the tree.
Kruskal’s algorithm is often not expressed through formal mathematical equations, but rather through an algorithmic description. Its logic is based on ordering of arcs and the management of connected components through union and search operations. Kruskal’s algorithm uses a greedy strategy that selects edges in ascending order with respect to weights, making it more efficient than a purely “brute force” approach.
Here is an overview of how Kruskal’s algorithm works:
- Initialization: Sort the edges of the graph in ascending order with respect to the weights.
- Initialize the tree: Start with a graph that consists only of the original nodes and no edges.
- Iterate over edges: Start adding edges to the graph, starting with those with the lowest weights. Make sure that adding an edge does not create a cycle in the current graph.
- Loops and Union-Find: To avoid the formation of loops, the algorithm uses a data structure called Union-Find (or Disjoint Set) to keep track of the connected components of the graph. Before adding an edge, check whether the nodes of that edge belong to the same component with Union-Find. If they don’t belong to the same component, add the arc and join the two components.
- Termination: Continue adding edges until all nodes are connected or until all edges are exhausted.
Kruskal’s algorithm is efficient and has a time complexity of ![]() , where E is the number of edges in the graph. It is a common choice when looking for an MST algorithm in a sparse graph or when dealing with large graphs.
, where E is the number of edges in the graph. It is a common choice when looking for an MST algorithm in a sparse graph or when dealing with large graphs.
There are other alternatives for finding a Minimum Spanning Tree, such as Prim’s algorithm. Prim’s algorithm uses a similar approach, but selects edges based on minimum node weights, rather than sorting all edges initially. The choice between Kruskal and Prim often depends on the graph structure and specific application requirements. Some optimizations, such as the use of binary heaps for edge sorting, can be applied to further improve the performance of Kruskal’s algorithm.
Graphs and the Networkx library
Kruskal’s algorithm can be used on undirected and unweighted graphs. In particular, it is suitable for undirected graphs (each edge has the same importance regardless of direction) and weighted graphs (each edge has an associated weight). The graph can be connected or unconnected, but its application is more common on connected graphs, as it tries to find a spanning tree that connects all nodes.
Now, let’s proceed with the graphical representation of the graph using the library
import networkx as nx
import matplotlib.pyplot as plt
# Example graph with 7 elements
graph = {
'A': [('B', 2), ('C', 1), ('D', 4)],
'B': [('A', 2), ('C', 3), ('D', 2)],
'C': [('A', 1), ('B', 3), ('D', 5)],
'D': [('A', 4), ('B', 2), ('C', 5)],
'E': [('D', 1)],
'F': [('A', 2), ('C', 4)],
'G': [('D', 3), ('F', 1)]
}
# Creation of the networkx graph
G = nx.Graph()
# Add nodes and edges to the networkx graph
for node in graph:
G.add_node(node)
for neighbor, weight in graph[node]:
G.add_edge(node, neighbor, weight=weight)
# Place nodes with a layout algorithm
pos = nx.spring_layout(G)
# Draw the graph
nx.draw(G, pos, with_labels=True, font_weight='bold', node_size=700, node_color='skyblue')
# Label the weights of the edges
labels = {(edge[0], edge[1]): edge[2]['weight'] for edge in G.edges(data=True)}
nx.draw_networkx_edge_labels(G, pos, edge_labels=labels)
plt.title("Example Graph with 7 Elements")
plt.show()
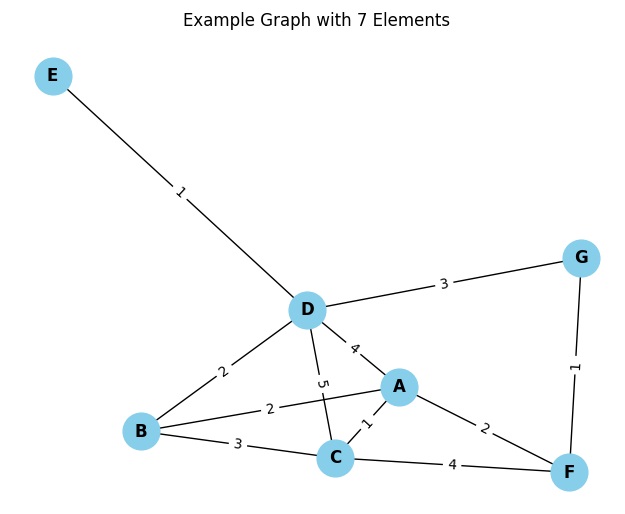
In this code, we are using the networkx library to create the graph and matplotlib for visualization. The nx.Graph() function creates a new empty graph, and then through a for loop, we add nodes and edges to the networkx graph. Finally, we use nx.draw() to draw the graph with node labels and nx.draw_networkx_edge_labels() to add labels to the edge weights. The plt.show() function displays the graph.
By executing you obtain the graphical representation of the graph in question.
MST (Minimum Spanning Tree)
MST (Minimum Spanning Tree), which translated into Italian means “Minimum Spanning Tree”. An MST of an undirected, weighted graph is a subset of the edges that connects all nodes of the graph without forming cycles and has the lowest possible total weight. In other words, it is a tree that connects all the nodes of a graph with the minimum possible cost.
Some key concepts associated with an MST are:
- Spanning Tree: A spanning tree of a graph is a subset of edges that forms a tree and touches all the nodes of the graph.
- Weighting: Each edge in a weighted graph has an associated value called “weight”. The goal is to find a subset of edges whose total weight is minimum.
MST is used in various contexts, including:
- Communication Networks: In telecommunications or computer networks, MST can be used to design low-cost connection networks between various points.
- Electrical Circuit Design: In electrical circuit design, MST can represent a low-cost connection between various components.
- Network Routing: In the design of computer or data transport networks, MST can be used to determine least-cost transmission paths.
- Data Clustering: In data analytics or machine learning, MST can be used to identify meaningful relationships and connections between data.
MST is a fundamental concept in graph theory and provides an optimal solution to minimum-cost connection problems in many practical applications. Algorithms such as Kruskal’s algorithm or Prim’s algorithm are often used to find the MST of a graph.
Application of Kruskal’s algorithm
Here is a version of the code that applies Kruskal’s algorithm to the example graph and provides a graphical representation:
import networkx as nx
import matplotlib.pyplot as plt
def find_mst_kruskal(graph):
edges = []
for node in graph:
for neighbor, weight in graph[node]:
edges.append((node, neighbor, weight))
edges.sort(key=lambda x: x[2])
mst_edges = []
parent = {node: node for node in graph}
def find(node):
if parent[node] != node:
parent[node] = find(parent[node])
return parent[node]
def union(node1, node2):
root1 = find(node1)
root2 = find(node2)
parent[root1] = root2
for edge in edges:
node1, node2, weight = edge
if find(node1) != find(node2):
mst_edges.append(edge)
union(node1, node2)
return mst_edges
# Example graph with 7 elements
graph = {
'A': [('B', 2), ('C', 1), ('D', 4)],
'B': [('A', 2), ('C', 3), ('D', 2)],
'C': [('A', 1), ('B', 3), ('D', 5)],
'D': [('A', 4), ('B', 2), ('C', 5)],
'E': [('D', 1)],
'F': [('A', 2), ('C', 4)],
'G': [('D', 3), ('F', 1)]
}
# Apply Kruskal's algorithm
minimum_spanning_tree = find_mst_kruskal(graph)
print("Minimum Spanning Tree:", minimum_spanning_tree)
# Creation of the networkx graph
G = nx.Graph()
# Add nodes and edges to the networkx graph
for node in graph:
G.add_node(node)
for neighbor, weight in graph[node]:
G.add_edge(node, neighbor, weight=weight)
# Place nodes with a layout algorithm
pos = nx.spring_layout(G)
# Draw the original graph
plt.figure(figsize=(12, 6))
plt.subplot(121)
nx.draw(G, pos, with_labels=True, font_weight='bold', node_size=700, node_color='skyblue')
plt.title("Original Graph")
# Draw the MST
plt.subplot(122)
nx.draw(G, pos, with_labels=True, font_weight='bold', node_size=700, node_color='skyblue')
nx.draw_networkx_edges(G, pos, edgelist=minimum_spanning_tree, edge_color='red', width=2)
plt.title("Minimum Spanning Tree")
plt.show()Explanation of the main parts of the code:
- find_mst_kruskal: This function implements Kruskal’s algorithm for finding the Minimum Spanning Tree. Use the Union-Find data structure to manage the connected components of the graph.
- Applying the Algorithm to an Example Graph: An example graph with 7 elements is created and Kruskal’s algorithm is applied to it to find the MST.
Creation of the networkx graph and graphical representation:
- A networkx graph is created and edges are added with their respective weights.
- The nx.spring_layout function is used to position nodes with a layout algorithm.
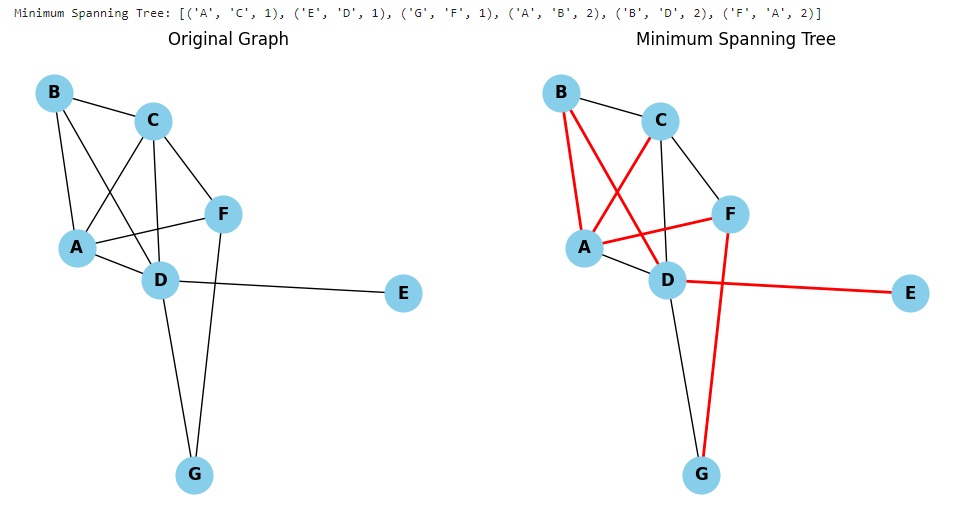
The graphical representation of the original graph and the resulting MST is shown in two separate subplots. MST arcs are highlighted in red.
Running the code you get:
Conclusion
In conclusion, Kruskal’s algorithm and the concepts associated with MST are fundamental in graph theory and have numerous practical applications, especially when it comes to minimum-cost connections in various domains. Graphing is a valuable tool for visually understanding relationships and structures in data.